|
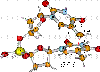
We consider diffusion-controlled reactions with applications mainly in
molecular biology. Since microscopic models of realistically large
reaction networks are prohibitively complex, well stirred chemically
reacting systems are often modeled by a continuous-time Markov chain
(chemical master equation).
For problems with spatial dependence, a suitable discrete version of
Brownian motion in the computational domain can be combined with well
stirred chemical kinetics to obtain the reaction-diffusion master
equation. This is a viable stochastic model at the mesoscopic
scale. It was previously not well understood how to sample from this
model other than for the simple case of Cartesian meshes.
In this talk I will discuss the conditions for the validity of this
approach and show how to evolve the model in a consistent way using
unstructured meshes. The resulting method is a flexible hybrid
algorithm in that the diffusion can be handled either in the
stochastic or in the deterministic domain. Possible applications of
the method are illustrated with examples inspired by molecular
biology.
|